Most people with Li Fraumeni Syndrome know they have a mutation. Some can even tell you at which codon their mutation occurred(don't feel bad if you had to look yours up-I had to). I'm a Gly245Ser- it's a hot spot binding domain within the tp53 tumor suppressor gene. I have an Adenine where I should have a Guanine which means the codon then codes for Serine instead of Glycine. Other hot spot codons are 175, 248, 249, 273 and 282. Any of this sound like Greek? We could all belong to a special LFS fraternity- except there is no L or F in the greek alphabet- go figure.
Did you ever wonder what p53 looks like? The book cover above shows what one version of p53 looks like- done by xray crystallography- a way of using xrays and the way the beams bend around atoms to "see" a 3 dimensional structure of these tiny tiny proteins.
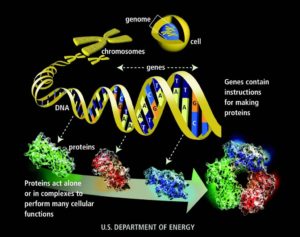
To have LFS, you inherited a mutated copy of the gene that makes the p53 protein. The p53 protein does many things in the body- but one of it's very important jobs is to keep tumor cells from going crazy. Sometimes, a person finds they are the first one in their family to have this unlucky mutation- they are de novo mutants- they did not inherit the faulty gene from a parent- but some environmental stress or other cause mutated their copy of TP53 along the way. They could pass along this to their children. The good news is we all have 2 copies of genes- one from our mom, one from our dad(or one newly mutated and one not mutated). So there is a chance we could pass along a healthy copy too. TP53 refers to the gene- pieces of DNA that are the blueprint to make proteins. p53 is the protein. The p53 protein can bind to other proteins- each different binding can change the shape- and giving the complex a different function. Sometimes, when there is a mutation- the protein cannot bind to certain proteins and it no longer works- it can no longer stop tumor cells from going haywire.
The gene TP53 has 393 amino acids in sequence that when put together make the entire protein. Each amino acid contains a set of 3 nucleotides. There are 4 nucleotides- they are the letters you see on pictures of DNA- AGCT(Adenine, Guanine,Cytosine and Thymine). A's like to hang with T's, C's like to hang with G's. They hold hands which then connects the 2 strands of DNA. Because, like us- these characters have dynamic personalities- they tend to arrange themselves near their friends- the other base pairs they are comfortable with- which cause the strands to coil and fold in a certain way. Remember playing the telephone game in class? Everyone lines up and the teacher gives the first student a phrase and she then relays that to the next student, who passes it along until it reaches the end. Sometimes the phrase is intact at the end, sometimes a student makes a mistake- and the end product may be different. The same thing happens in DNA when it unfolds to make it's message(the protein). The message can get mixed up. Sometimes the message still makes sense, sometimes it is completely confusing. When mutations happen- sometimes the cells can still make the protein- it only works a little bit. But as the cell gets older, more and more mistakes are made and eventually it doesn't work at all. So think about all our friends playing the telephone game- stretching out in a line, holding hands to relay the message- when the message is done- they still have to hold hands- but want to talk with their other friends- they have to bend and move in a way to get there. That is how proteins fold up to make their shapes.
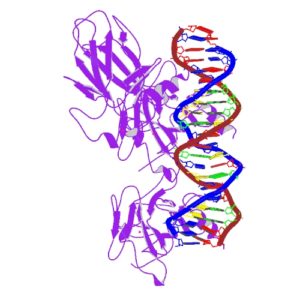 |
| IARC image of p53 bound to DNA |
There is a set order of nucleotides(ACGT) that code for a gene. When mutations happen, this set order changes. 3 nucleotides make an amino acid and fortunately sometimes 3 different nucleotides can make the same amino acid- so if this mutation occurs- it may not be as bad. It's like having 3 foods on your plate- you can choose the order you eat them in- sometimes things taste funky together and sometimes they pair well. If they pair well it doesn't always matter which one you eat first. Let's think of the tp53 gene like a zipper. There is a set order and the friends match up with friends- holding hands to keep everything together. Each tooth of the zipper is one nucleotide. It takes 3 teeth to make an amino acid. If you take out a tooth(a mutation type called a deletion) the zipper will get stuck. Sometimes depending on where this is at, you could shift down a bit and still make it zip - but it will be off. Sometimes a tooth can be replaced where there is a hole(called a point mutation) and can have one of 3 effects. If it is the same type of tooth- the zipper will work(silent mutation- the nucleotide codes for the same amino acid in the end). If the tooth is different- the zipper may stick a little but still work(missense mutation-the nucleotide will change the code to a different amino acid). If the tooth is too different- it will break the zipper(nonsense mutation- no amino acid can be made.)
Here's an example of our zipper effects(mutations);
the cat ate the rat on the mat
Each word is a codon.
thecatatetheratonthemat
If we delete the 5th nucleotide we are left with:
thectatetheratonthemat
the cta tet her ato nth ema -the whole sentence is affected.
If we insert another nucleotide:
thecotatetheratonthemat
the cot ate the rat on the mat - one codon is different, and even though the majority of the sentence still makes sense- that one codon difference can change the meaning of the sentence.
 |
| IARC table of tp53 sequence |
With 393 amino acids, that means there are over 1100 nucleotides that could be mutated. Some spots are more exposed than others or more susceptible to damage. Sometimes a break at the beginning or the end of the zipper means it won't work, whereas you can gloss over one in the middle. This is one reason why it's so tough to link specific cancers to specific mutations. We are rare breed of rare cancers and rare mutations. In Brazil- there was a huge step forward when researchers discovered that a particular mutation of the 337 codon could be linked to adrenocortical carcinoma(ACC). Unfortunately there aren't always clusters of information like this to help scientists find the links. In Brazil they found 72 families with this rare mutation, also linked to a core LFS cancer- yet these families do not have LFS. As a result, they instituted a newborn screening program to help catch these children and monitor them for ACC to catch and treat it at the earliest stages.
The hope is that in the future we can not only identify all of the specific p53 mutations of LFS, but to provide working therapy to either help the protein work again or prevent further damage. The other avenue is to identify a link between mutations and cancer type to narrow down the screening and most beneficial therapies tailored to specific mutants. Ultimately gene therapy could hopefully someday provide a way to fix these mutations.
Resources
IARC TP53 Database -p53.iarc.fr
TP53 sequence
Mutation Notation IARC p53
Clinical Outcomes and Correlates of TP53 mutations and cancer